Streamlined Tandem Mass Tag (SL-TMT) Protocol: An Efficient Strategy for Quantitative (Phospho)proteome Profiling Using Tandem Mass Tag-Synchronous Precursor Selection-MS3. - Abstract - Europe PMC

Sample Preparation for Relative Quantitation of Proteins Using Tandem Mass Tags (TMT) and Mass Spectrometry (MS). - Abstract - Europe PMC

Streamlined Tandem Mass Tag (SL-TMT) Protocol: An Efficient Strategy for Quantitative (Phospho)proteome Profiling Using Tandem Mass Tag-Synchronous Precursor Selection-MS3. - Abstract - Europe PMC

AminoxyTMT: A novel multi-functional reagent for characterization of protein carbonylation | BioTechniques
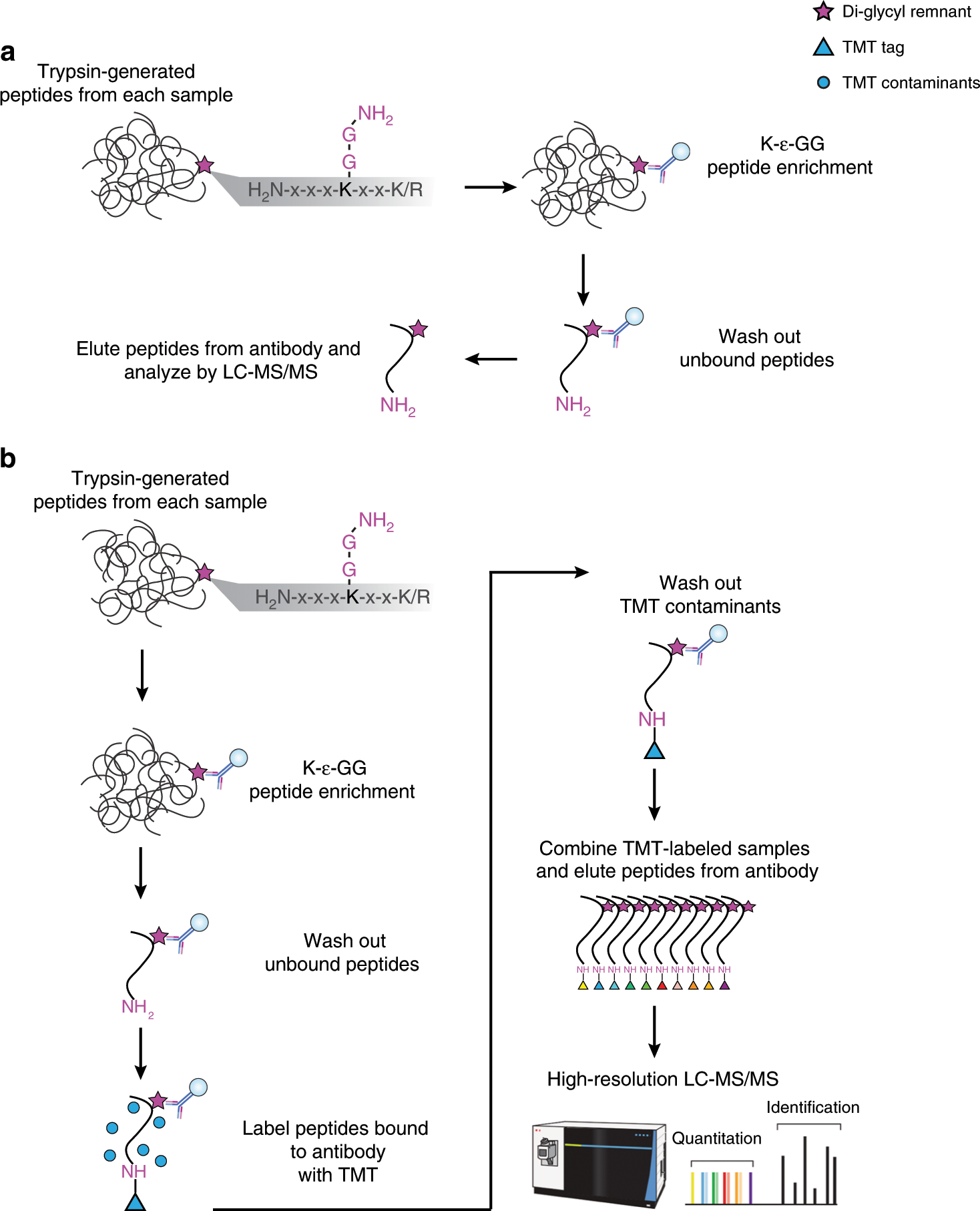
Rapid and deep-scale ubiquitylation profiling for biology and translational research | Nature Communications

Applications of iTRAQ and TMT Labeling Techniques to the Study of Neurodegenerative Diseases | Bentham Science

High-throughput and Deep-proteome Profiling by 16-plex Tandem Mass Tag Labeling Coupled with Two-dimensional Chromatography and Mass Spectrometry | Protocol

Multibatch TMT Reveals False Positives, Batch Effects and Missing Values* - Molecular & Cellular Proteomics

Extended Multiplexing of Tandem Mass Tags (TMT) Labeling Reveals Age and High Fat Diet Specific Proteome Changes in Mouse Epididymal Adipose Tissue* - Molecular & Cellular Proteomics
News in Proteomics Research: TMT Labeling for the Masses! A thought experiment for ultra-efficient, low cost labeling!

Experimental design and TMT labelling workflow for the experiment.: (A)... | Download Scientific Diagram
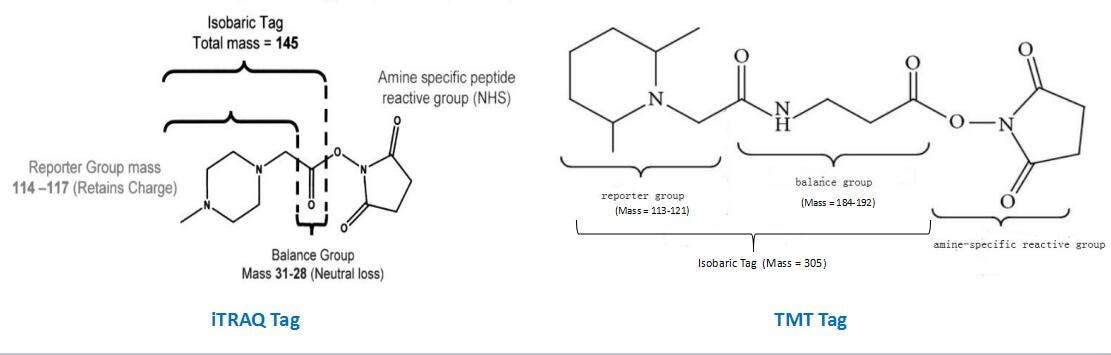
Part I: Detailed Description of iTRAQ/TMT Tag Structure and Relative Quantification Principle | by Prime Jones | Medium

Deep Proteome Profiling by Isobaric Labeling, Extensive Liquid Chromatography, Mass Spectrometry, and Software-assisted Quantification | Protocol

High-Throughput Profiling of ProteomeModifications and Posttranslational by 16-Plex TMT Labeling and Mass Spectrometry | SpringerLink